Differential Inhibition of mRNA Degradation Pathways by Novel Cap Analogs* - Journal of Biological Chemistry

A Mammalian Pre-mRNA 5′ End Capping Quality Control Mechanism and an Unexpected Link of Capping to Pre-mRNA Processing - ScienceDirect

Rapid Analysis of Synthetic mRNA Cap Structure Using Ion-Pairing RPLC with the BioAccord LC-MS System | Waters

CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology

CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology
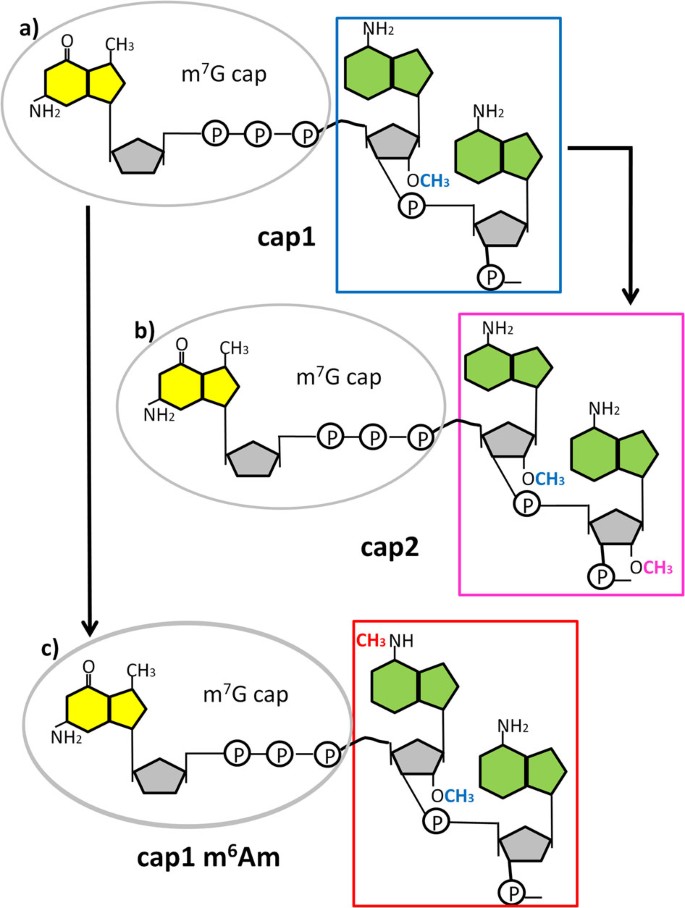
A novel synthesis and detection method for cap-associated adenosine modifications in mouse mRNA | Scientific Reports